I am going to apply the same principle on DeSEQ as well. I will be using the Benjamini-Hochberg false discovery rate of 0.01 for EdgeR based on a recent paper. On top of this, I am thinking of doing a normalization of the counts to see how the data appears in normalized data sets. Cuffdiff already does a statistical test via its algorithm. I am thinking of doing a negative binomial test as I am comparing the expression presence between two organs. Thank you!I am currently debating, which statistical tests will be best to use. 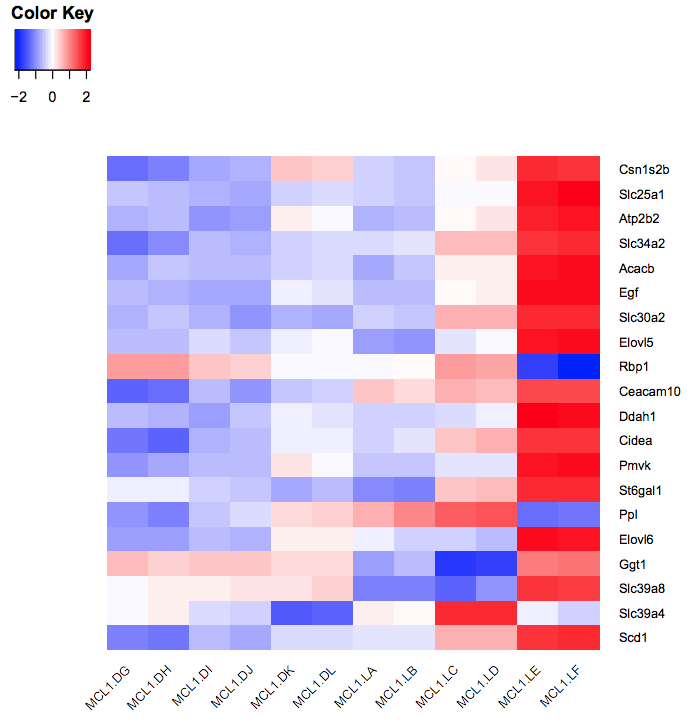
How Zapages How do you compare your DE result from DEseq/edgeR/cuffdiff? By which statistical variable? I am a rookie in RNA-seq. Is the binomial test fine for DESeq? Can that be then be used for creating a bar/histogram? Any statistical test that you would recommend for Graphpad or through DESeq/EdgeR? I don't think graph pad will allow me to create bar graphs with error bars unless I do a statistical test. Although Cummerbund is fairly easy to use to make these graphs for the tuxedo pipeline. Should I just use the normalized values for the genes in Graphpad? I am learning R, any advice on creating these graphs with error bars. Yes its true, we are hoping to publish this data.Īs for creating bar graphs. Gene ID | locus | sample_1 Name | sample_2 Name| status | value_1 |value_2 log2(fold_change) |test_stat| p_value |q_value | significant (Yes/No) The gene_exp.diff file has the following information: Gene ID|baseMean|Base Mean Sample 1|Base Mean Sample 2| Normalized value|log2 fold-change | p value | q value | So something like this should be fine for both DESeq and EdgeR tables: Thank you dpryan for all the great advice. If someone could kindly help me on how I could go about this. In total I hope to have 3 Heatmaps with the specific genes of interests only: The p values and log information seem to indicate high values for the genes that I am interested in. I have 2 replicates for Organ A and 2 replicates for Organ B. I have already used Tuxedo pipeline to create a heat map using CummeRbund for the species. Or is there any better method to analyze these genes via EdgeR and DeSeq. Now I was wondering how could I possibly create heat map based on the genes that I interested based on EdgeR and DeSeq. Edited/removed no_feature, ambiguous, too_low_aQual:, not_aligned, alignment_not_unique in the output file and added Organs headers for each rep.ĥ) Used EdgeR and DeSeq respectively on Iplant Collaborative. (offline)Ĥ) Used Iplant Collaborative Join tab delimited to merge the Counts into one file. I have done the following:ġ) Tophat2 (provided Ensembl GTF and Fasta files)Ģ) Used Samtools to convert the files to Sam format (offline)ģ) HTseq and used the same GTF file from Tophat2. This X-Function is used for importing Prism (PZFX, XML) files.I am new with DeSeq and EdgeR. This is used to set the import options in the dialog. It allows to select which tables in the file to be imported. This is used to show the file information. In the dialog, click the Browse button beside the list box to open one or more files and the filenames will be listed in the box. Specify the filename(s) of the file(s) to be imported. Please refer to the page for additional option switches when accessing the x-function from script Variables Display Minimum Origin Version Required: 8.5.1SR0 To add drag & drop support for a file type, see Import Filter Manager. If you do not see this file type, choose Data: Import from File: Add/Remove File Types. Data: Import from File: Prism (PZFX, XML)
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
